Difference between revisions of "Dynamo Workshop at the International School of Crystallography 2019"
Jump to navigation
Jump to search
| Line 3: | Line 3: | ||
== Computers == | == Computers == | ||
| + | |||
=== Download === | === Download === | ||
| − | + | The Dynamo versions on the three platforms can be obtained in the [[Downloads]] page. Please use the indications [[Installation|here]] to install it in your laptop. | |
=== Starting a ''Dynamo'' session === | === Starting a ''Dynamo'' session === | ||
| − | + | ||
== Program == | == Program == | ||
Revision as of 08:12, 5 June 2019
This page describes the contents of the practical hands-on sessions for tomography through Dynamo in at the 54th International School of Crystallography in Erice. A core presentation will be held on site
Contents
Computers
Download
The Dynamo versions on the three platforms can be obtained in the Downloads page. Please use the indications here to install it in your laptop.
Starting a Dynamo session
Program
Before the hands-on session, we will go to this presentation for a general introduction of the Dynamosoftware.
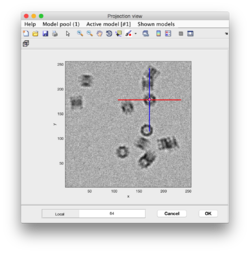
General Introduction

Clicking particles in the Starters guide
Guided presentation:
- tutorial on basic elements: help, data and metadata formats.
- tutorial on the basic concept in Dynamo alignment: the project.
Working on your own:
- Basic walkthrough: creating a catalogue, picking particles, launching a project.
- Advanced starters guide on a real tomogram. (~2 hours)
- Further work:
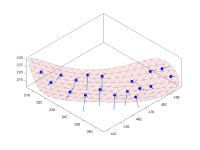
Geometric modeling
Short guided presentation:
- tutorial on membrane modeling with dmslice
- Filament models with dtmslice
- Reusing model workflows ( walkthrough)
- Further work: catalogue
Working on your own:
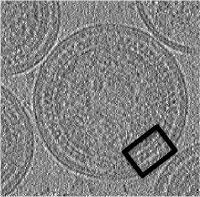
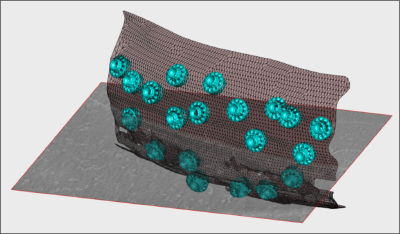
- In the afternoon, after the research talks, we will focus on the extraction of particles from densely packed spherical geometry on HIV viral capsides (~1 hour)
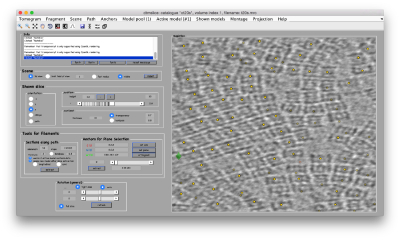
Template matching
Working on your own:
- We will follow this walkthrough for automated identification of proteosomes on a real tomogram through template matching. (~1 hour)
Adaptive bandpass filtering
Working on your own:
- We will follow this walkthrough to create a small synthetic data set that illustrates the principles of adaptive bandpass filtering, a way of conducting a golden standard alignment procedure . (~40 mins)
Classification
Short guided presentation:
- PCA Basic concepts.
advanced subboxing tutorial: combining with PCA and MRA tutorial - Multirefererence Analysis
Basic concepts.
walkthrough (~10 min) - Further work
- Commandline operations with PCA
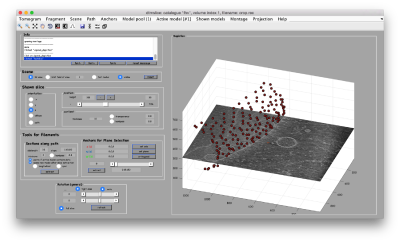
Creation of 3D scenes
Working on your own:
- Walkthrough on the FHV data set (~1hour)
Further support material.
- Walkthrough on depiction and manipulation of triangulations (synthetic data).
Additional tools
Wednesday afternoon session.
- Subboxing
- motivation slide: extraction of vertices from icosahedral viruses
- Basic subboxing (tutoriall)
- advanced subboxing tutorial: combining with MRA tutorial
- suggested data set: prd1 Capsides (in the folder of isolated particles)
- Manual alignment.
- Exercises
Data sets
The data sets will be updated here.
Instructors
- Daniel Castaño-Díez, University of Basel.